Large-scale multivariate sparse regression with applications to UK Biobank
Jul 19, 2022
·
1 min read
Abstract
In high-dimensional regression problems, often a relatively small subset of the features are relevant for predicting the outcome, and methods that impose sparsity on the solution are popular. When multiple correlated outcomes are available (multitask), reduced rank regression is an effective way to borrow strength and capture latent structures that underlie the data.
In high-dimensional regression problems, often a relatively small subset of the features
are relevant for predicting the outcome, and methods that impose sparsity on the
solution are popular. When multiple correlated outcomes are available (multitask),
reduced rank regression is an effective way to borrow strength and capture latent
structures that underlie the data. Our proposal is motivated by the UK Biobank
population-based cohort study, where we are faced with large-scale,
ultrahigh-dimensional features, and have access to a large number of outcomes
(phenotypes)-lifestyle measures, biomarkers, and disease outcomes. We are hence led to
fit sparse reduced-rank regression models, using computational strategies that allow us
to scale to problems of this size. We use a scheme that alternates between solving the
sparse regression problem and solving the reduced rank decomposition. For the sparse
regression component we propose a scalable iterative algorithm based on adaptive
screening that leverages the sparsity assumption and enables us to focus on solving much
smaller subproblems. The full solution is reconstructed and tested via an optimality
condition to make sure it is a valid solution for the original problem. We further
extend the method to cope with practical issues, such as the inclusion of confounding
variables and imputation of missing values among the phenotypes. Experiments on both
synthetic data and the UK Biobank data demonstrate the effectiveness of the method and
the algorithm. We present multiSnpnet package, available at
http://github.com/junyangq/multiSnpnet that works on top of PLINK2 files, which we
anticipate to be a valuable tool for generating polygenic risk scores from human genetic
studies.
Type
Publication
Published in Ann. Appl. Stat., 2022
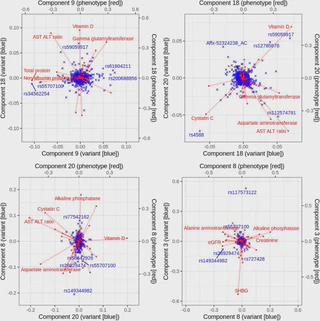
In this study led by Junyang Qian, we present a method to fit sparse multi-variate and multi-response regression model.