Significant Sparse Polygenic Risk Scores across 813 traits in UK Biobank
Abstract
We present a systematic assessment of polygenic risk score (PRS) prediction across more than 1,500 traits using genetic and phenotype data in the UK Biobank. We report 813 sparse PRS models with significant (p < 2.
We present a systematic assessment of polygenic risk score (PRS) prediction across more
than 1,500 traits using genetic and phenotype data in the UK Biobank. We report 813
sparse PRS models with significant (p < 2.5 x 10-5) incremental predictive performance
when compared against the covariate-only model that considers age, sex, types of
genotyping arrays, and the principal component loadings of genotypes. We report a
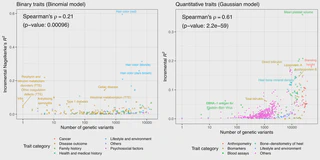
significant correlation between the number of genetic variants selected in the sparse
PRS model and the incremental predictive performance (Spearman’s ⍴ = 0.61, p = 2.2 x
10-59 for quantitative traits, ⍴ = 0.21, p = 9.6 x 10-4 for binary traits). The sparse
PRS model trained on European individuals showed limited transferability when evaluated
on non-European individuals in the UK Biobank. We provide the PRS model weights on the
Global Biobank Engine (https://biobankengine.stanford.edu/prs).
Type
Publication
Published in PLOS Genetics, 2022
We performed a systematic assessment of the predictive performance of PRS models across >1,500 traits in UK Biobank and report 813 PRS models with significant predictive performance.