Deconvoluting complex correlates of COVID19 severity with a multi-omic pandemic tracking strategy
Aug 30, 2022
·
0 min read
Abstract
The SARS-CoV-2 pandemic has differentially impacted populations across race and ethnicity. A multi-omic approach represents a powerful tool to examine risk across multi-ancestry genomes.
The SARS-CoV-2 pandemic has differentially impacted populations across race and
ethnicity. A multi-omic approach represents a powerful tool to examine risk across
multi-ancestry genomes. We leverage a pandemic tracking strategy in which we sequence
viral and host genomes and transcriptomes from nasopharyngeal swabs of 1049 individuals
(736 SARS-CoV-2 positive and 313 SARS-CoV-2 negative) and integrate them with digital
phenotypes from electronic health records from a diverse catchment area in Northern
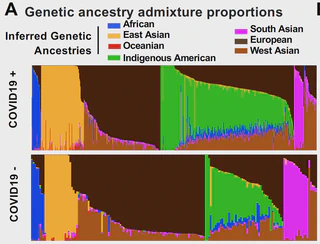
California. Genome-wide association disaggregated by admixture mapping reveals novel
COVID-19-severity-associated regions containing previously reported markers of
neurologic, pulmonary and viral disease susceptibility. Phylodynamic tracking of
consensus viral genomes reveals no association with disease severity or inferred
ancestry. Summary data from multiomic investigation reveals metagenomic and HLA
associations with severe COVID-19. The wealth of data available from residual
nasopharyngeal swabs in combination with clinical data abstracted automatically at scale
highlights a powerful strategy for pandemic tracking, and reveals distinct
epidemiologic, genetic, and biological associations for those at the highest risk.
Type
Publication
Published in Nat Commun., 2022