Bayesian model comparison for rare-variant association studies of multiple phenotypes
Nov 25, 2021
·
1 min read
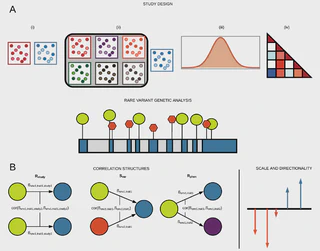
Abstract
Whole-genome sequencing studies applied to large populations or biobanks with extensive phenotyping raise new analytic challenges. The need to consider many variants at a locus or group of genes simultaneously and the potential to study many correlated phenotypes with shared genetic architecture provide opportunities for discovery not addressed by the traditional one variant, one phenotype association study.
Whole-genome sequencing studies applied to large populations or biobanks with extensive
phenotyping raise new analytic challenges. The need to consider many variants at a locus
or group of genes simultaneously and the potential to study many correlated phenotypes
with shared genetic architecture provide opportunities for discovery not addressed by
the traditional one variant, one phenotype association study. Here, we introduce a
Bayesian model comparison approach called MRP (multiple rare variants and phenotypes)
for rare-variant association studies that considers correlation, scale, and direction of
genetic effects across a group of genetic variants, phenotypes, and studies, requiring
only summary statistic data. We apply our method to exome sequencing data (n = 184,698)
across 2,019 traits from the UK Biobank, aggregating signals in genes. MRP demonstrates
an ability to recover signals such as associations between PCSK9 and LDL cholesterol
levels. We additionally find MRP effective in conducting meta-analyses in exome data.
Non-biomarker findings include associations between MC1R and red hair color and skin
color, IL17RA and monocyte count, and IQGAP2 and mean platelet volume. Finally, we apply
MRP in a multi-phenotype setting; after clustering the 35 biomarker phenotypes based on
genetic correlation estimates, we find that joint analysis of these phenotypes results
in substantial power gains for gene-trait associations, such as in TNFRSF13B in one of
the clusters containing diabetes- and lipid-related traits. Overall, we show that the
MRP model comparison approach improves upon useful features from widely used
meta-analysis approaches for rare-variant association analyses and prioritizes
protective modifiers of disease risk.
Type
Publication
Published in Am J Hum Genet, 2021
When applied to cardiometabolic biomarker traits in UK Biobank, MRP identified gene-biomarker associations that were not identified in single-variant GWAS analysis.