Polygenic risk modeling with latent trait-related genetic components
Feb 8, 2021
·
1 min read
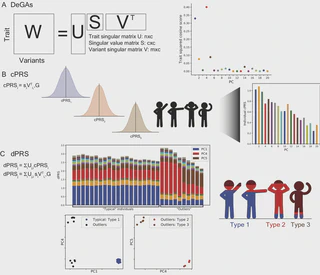
Abstract
Polygenic risk models have led to significant advances in understanding complex diseases and their clinical presentation. While polygenic risk scores (PRS) can effectively predict outcomes, they do not generally account for disease subtypes or pathways which underlie within-trait diversity.
Polygenic risk models have led to significant advances in understanding complex diseases
and their clinical presentation. While polygenic risk scores (PRS) can effectively
predict outcomes, they do not generally account for disease subtypes or pathways which
underlie within-trait diversity. Here, we introduce a latent factor model of genetic
risk based on components from Decomposition of Genetic Associations (DeGAs), which we
call the DeGAs polygenic risk score (dPRS). We compute DeGAs using genetic associations
for 977 traits and find that dPRS performs comparably to standard PRS while offering
greater interpretability. We show how to decompose an individual’s genetic risk for a
trait across DeGAs components, with examples for body mass index (BMI) and myocardial
infarction (heart attack) in 337,151 white British individuals in the UK Biobank, with
replication in a further set of 25,486 non-British white individuals. We find that BMI
polygenic risk factorizes into components related to fat-free mass, fat mass, and
overall health indicators like physical activity. Most individuals with high dPRS for
BMI have strong contributions from both a fat-mass component and a fat-free mass
component, whereas a few “outlier” individuals have strong contributions from only one
of the two components. Overall, our method enables fine-scale interpretation of the
drivers of genetic risk for complex traits.
Type
Publication
Published in European Journal of Human Genetics, 2021
Polygenic risk score (PRS) has been proposed for disease risk prediction with potential clinical relevance for some traits, but its personalized interpretation is generally difficult, especially when there exist disease subtypes driven by different genetic components.