Fast Lasso method for large-scale and ultrahigh-dimensional Cox model with applications to UK Biobank
Sep 29, 2020
·
1 min read
Abstract
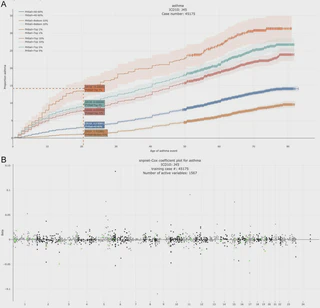
We develop a scalable and highly efficient algorithm to fit a Cox proportional hazard model by maximizing the $L^1$-regularized (Lasso) partial likelihood function, based on the Batch Screening Iterative Lasso (BASIL) method developed in Qian and others (2019). Our algorithm is particularly suitable for large-scale and high-dimensional data that do not fit in the memory.
We develop a scalable and highly efficient algorithm to fit a Cox proportional hazard
model by maximizing the $L^1$-regularized (Lasso) partial likelihood function, based on
the Batch Screening Iterative Lasso (BASIL) method developed in Qian and others (2019).
Our algorithm is particularly suitable for large-scale and high-dimensional data that do
not fit in the memory. The output of our algorithm is the full Lasso path, the parameter
estimates at all predefined regularization parameters, as well as their validation
accuracy measured using the concordance index (C-index) or the validation deviance. To
demonstrate the effectiveness of our algorithm, we analyze a large genotype-survival
time dataset across 306 disease outcomes from the UK Biobank (Sudlow and others, 2015).
We provide a publicly available implementation of the proposed approach for genetics
data on top of the PLINK2 package and name it snpnet-Cox.
Type
Publication
Published in Biostatistics, 2020
We propose extending the BASIL/snpnet algorithm to fit the L1 penalized Cox proportional hazards model using a large-scale dataset from a genotyped cohort.