Assessing Digital Phenotyping to Enhance Genetic Studies of Human Diseases
May 7, 2020
·
1 min read
Abstract
Population-scale biobanks that combine genetic data and high-dimensional phenotyping for a large number of participants provide an exciting opportunity to perform genome-wide association studies (GWAS) to identify genetic variants associated with diverse quantitative traits and diseases. A major challenge for GWAS in population biobanks is ascertaining disease cases from heterogeneous data sources such as hospital records, digital questionnaire responses, or interviews.
Population-scale biobanks that combine genetic data and high-dimensional phenotyping for
a large number of participants provide an exciting opportunity to perform genome-wide
association studies (GWAS) to identify genetic variants associated with diverse
quantitative traits and diseases. A major challenge for GWAS in population biobanks is
ascertaining disease cases from heterogeneous data sources such as hospital records,
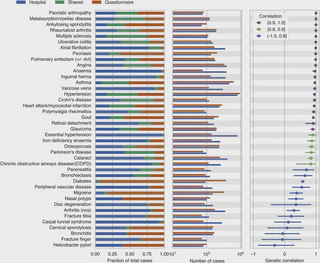
digital questionnaire responses, or interviews. In this study, we use genetic
parameters, including genetic correlation, to evaluate whether GWAS performed using
cases in the UK Biobank ascertained from hospital records, questionnaire responses, and
family history of disease implicate similar disease genetics across a range of effect
sizes. We find that hospital record and questionnaire GWAS largely identify similar
genetic effects for many complex phenotypes and that combining together both phenotyping
methods improves power to detect genetic associations. We also show that family history
GWAS using cases ascertained on family history of disease agrees with combined hospital
record and questionnaire GWAS and that family history GWAS has better power to detect
genetic associations for some phenotypes. Overall, this work demonstrates that digital
phenotyping and unstructured phenotype data can be combined with structured data such as
hospital records to identify cases for GWAS in biobanks and improve the ability of such
studies to identify genetic associations.
Type
Publication
Published in The American Journal of Human Genetics, 2020
Large-scale population-based genotyped biobanks with dense phenotypic information provide opportunities for genetic analysis at scale.