Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology
 Image credit: Adobe Stock
Image credit: Adobe StockPopulation-based biobanks with genomic and dense phenotype data provide opportunities for generating effective therapeutic hypotheses and understanding the genomic role in disease predisposition. To characterize latent components of genetic associations, we apply truncated singular value decomposition (DeGAs) to matrices of summary statistics derived from genome-wide association analyses across 2,138 phenotypes measured in 337,199 White British individuals in the UK Biobank study.
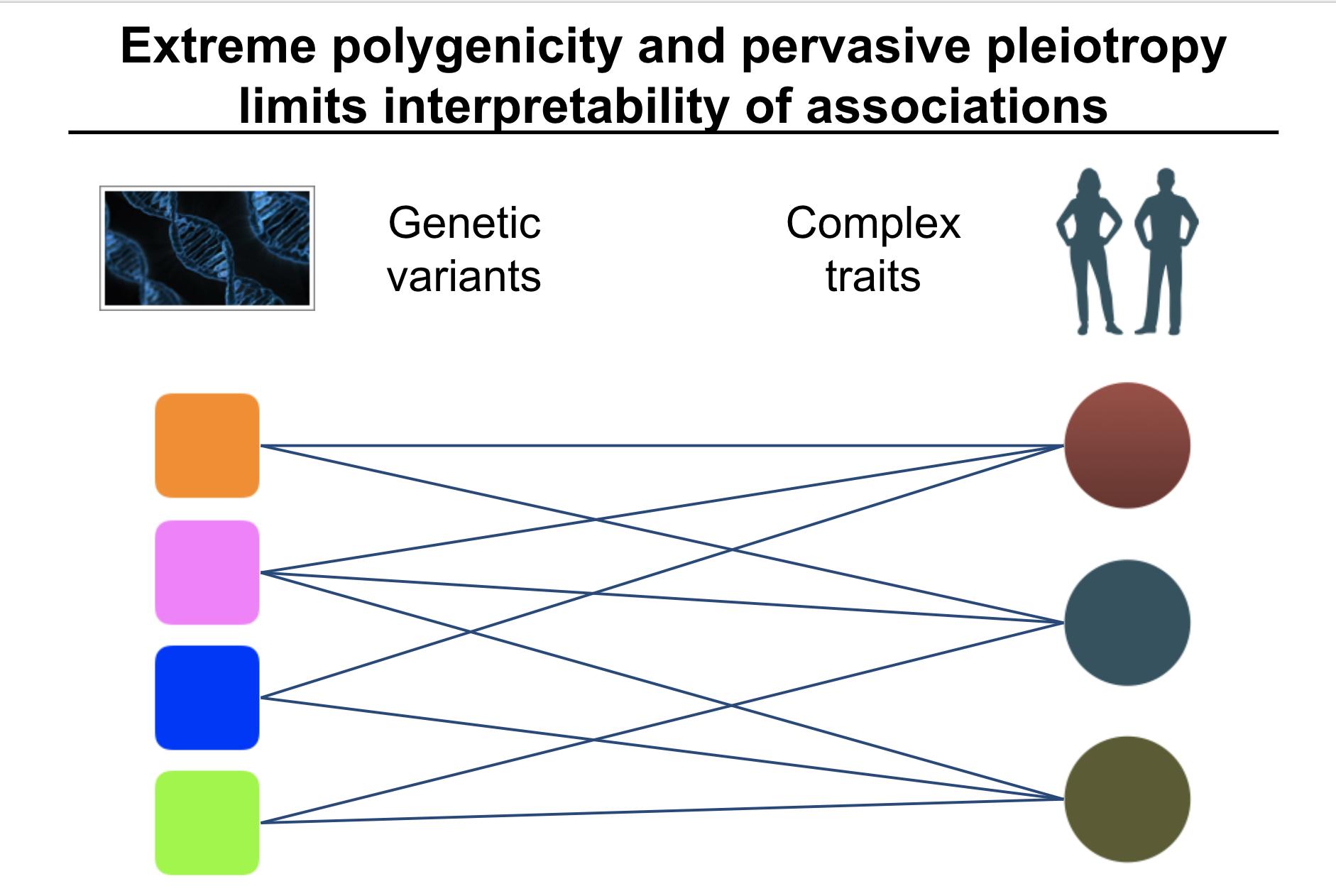
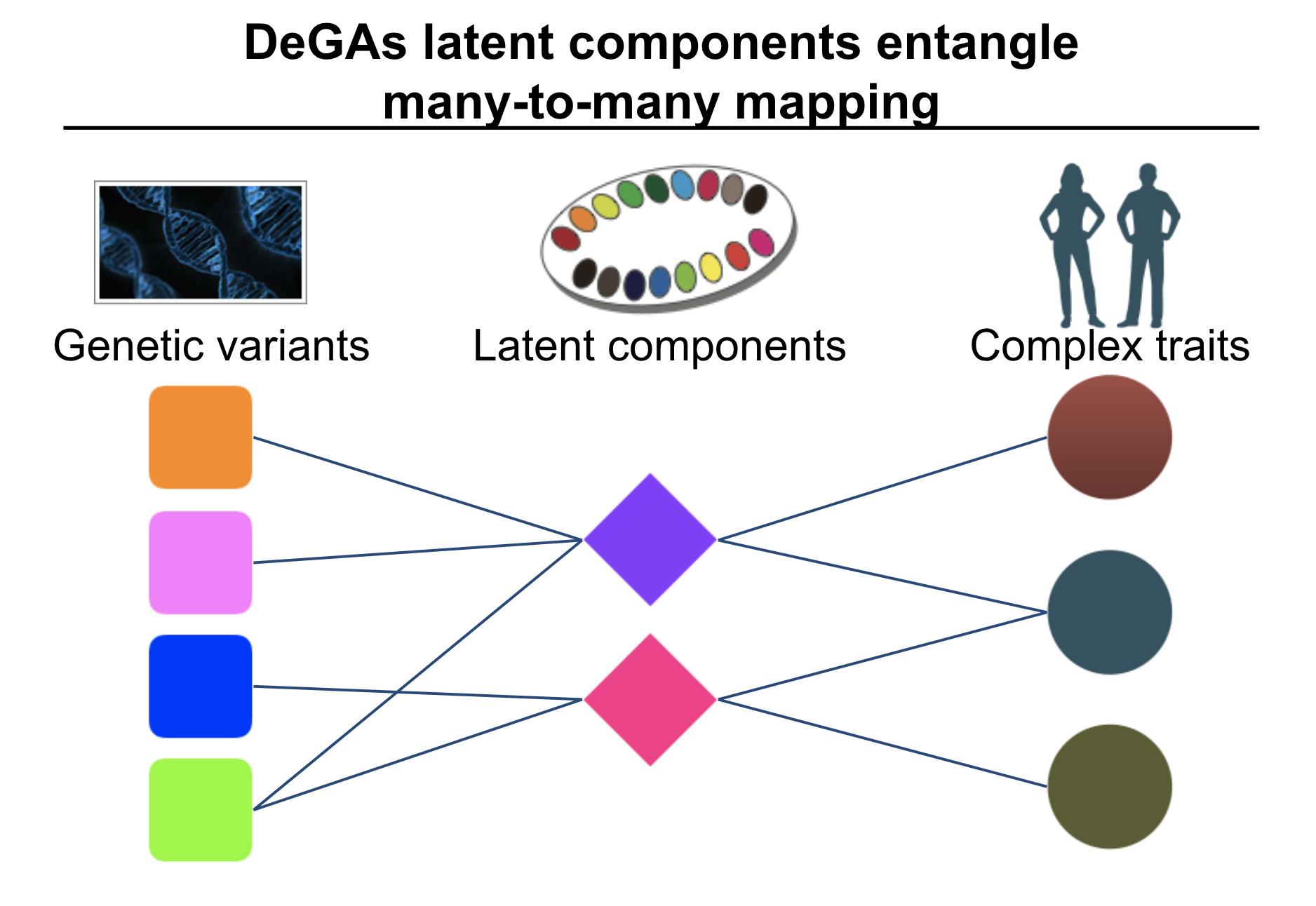
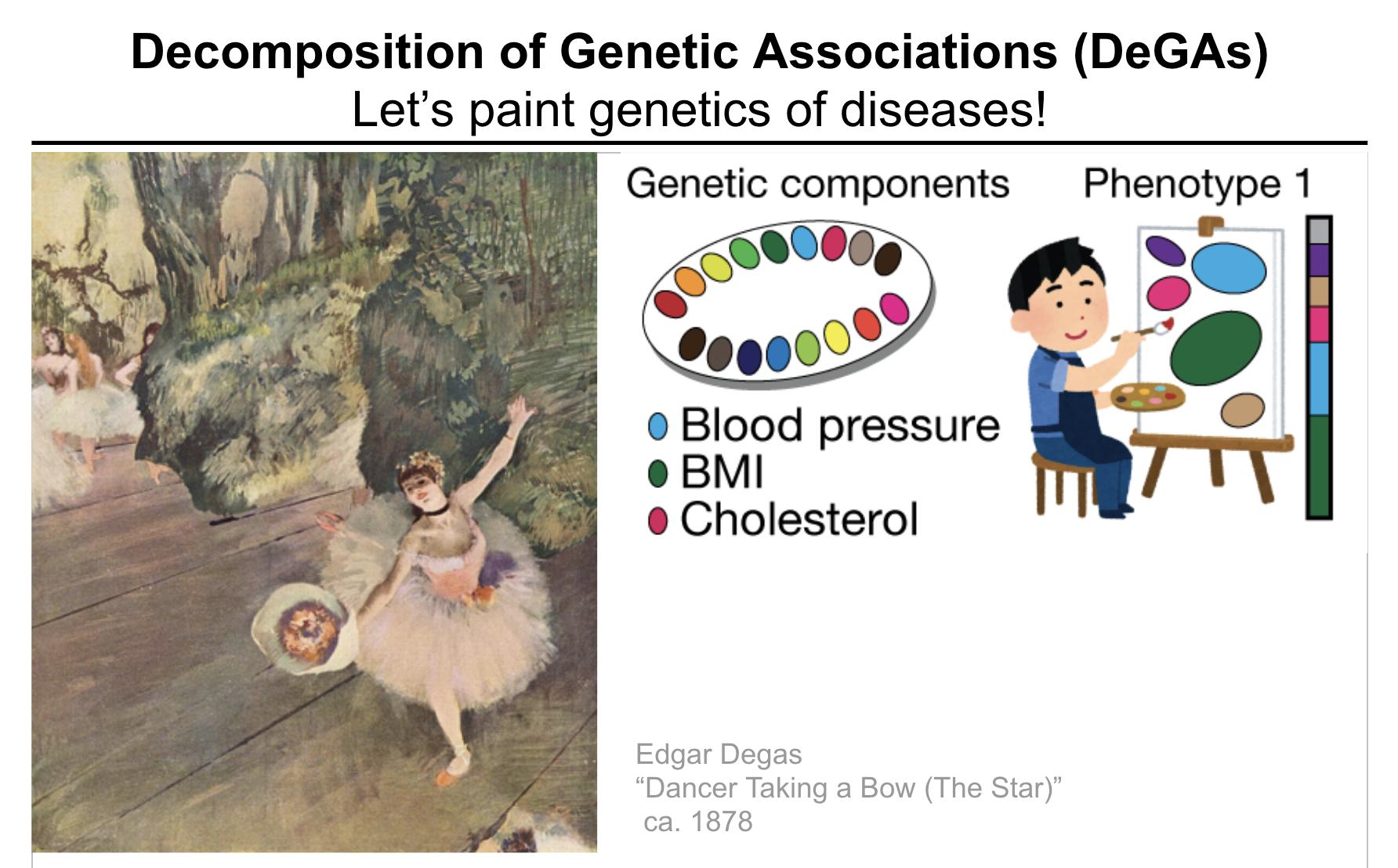
While many pleiotropic genetic loci have been identified, how they contribute to phenotypes across traits and diseases is unclear. We developed DeGAs to address this issue.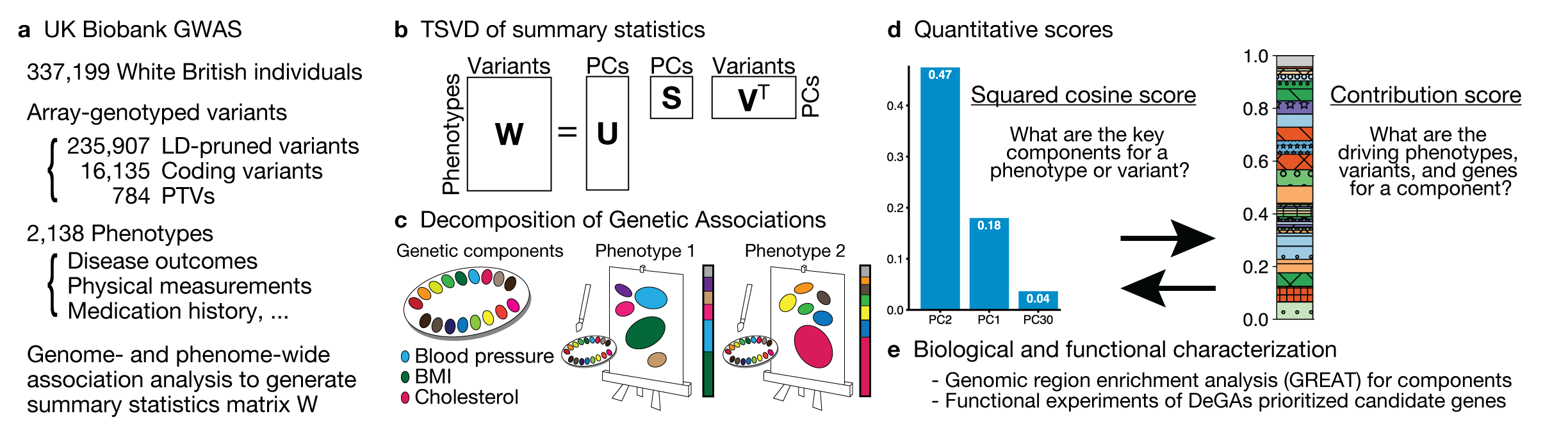
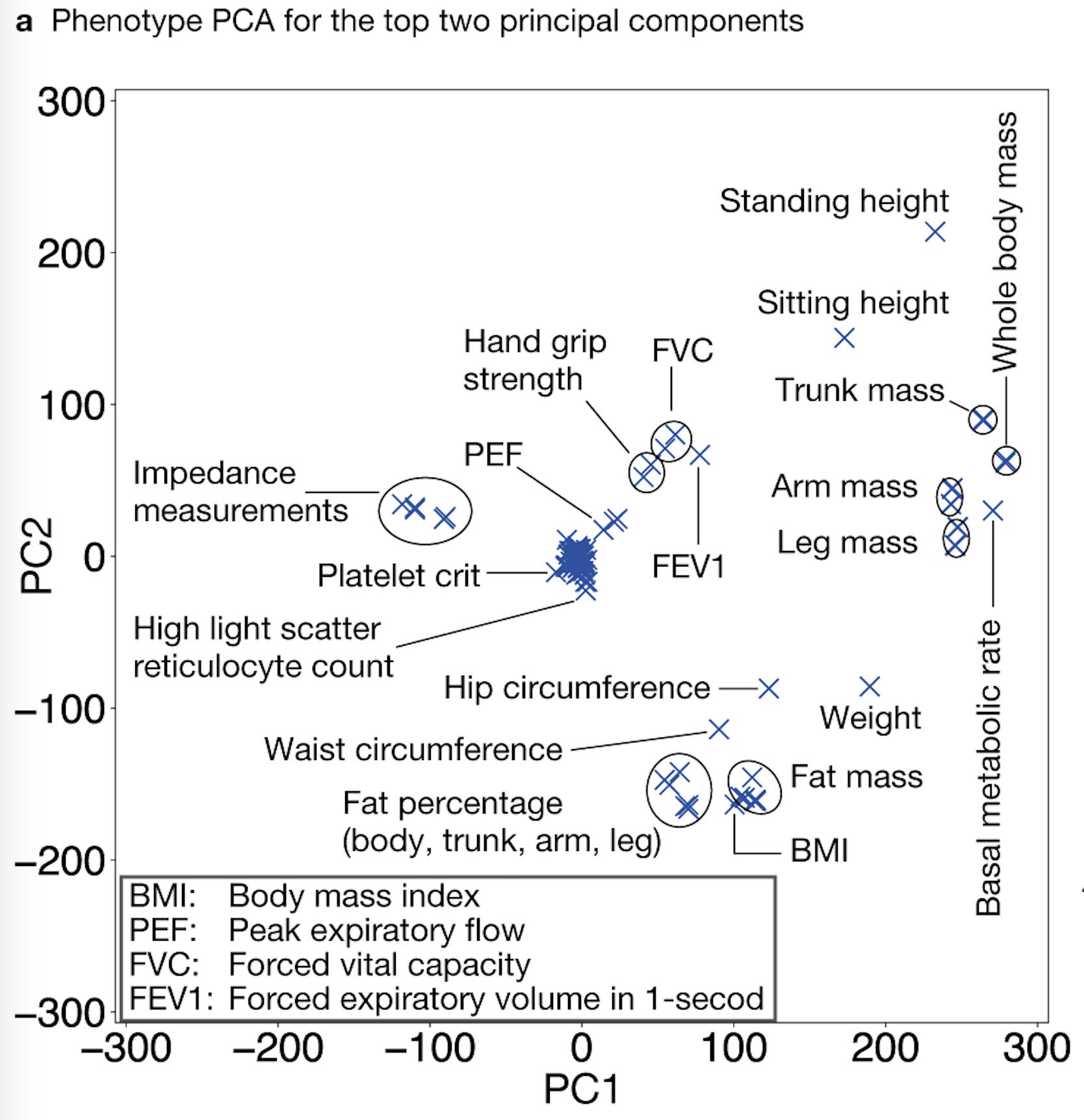
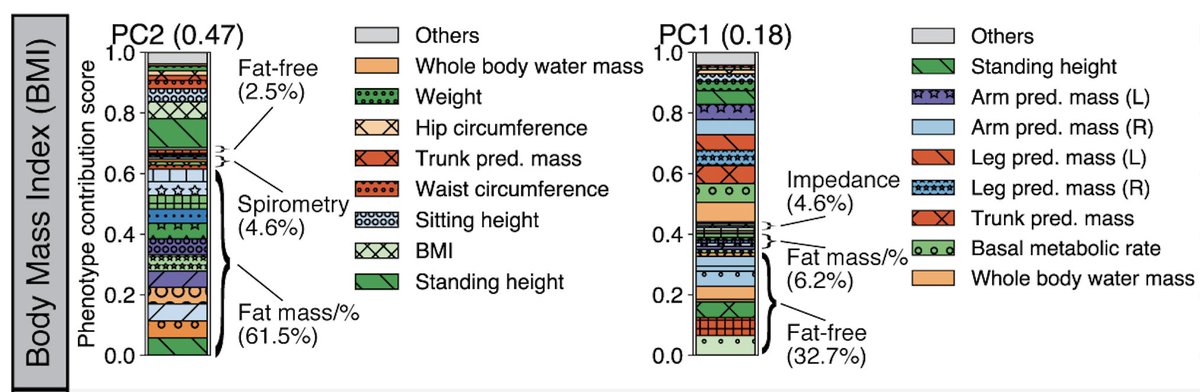
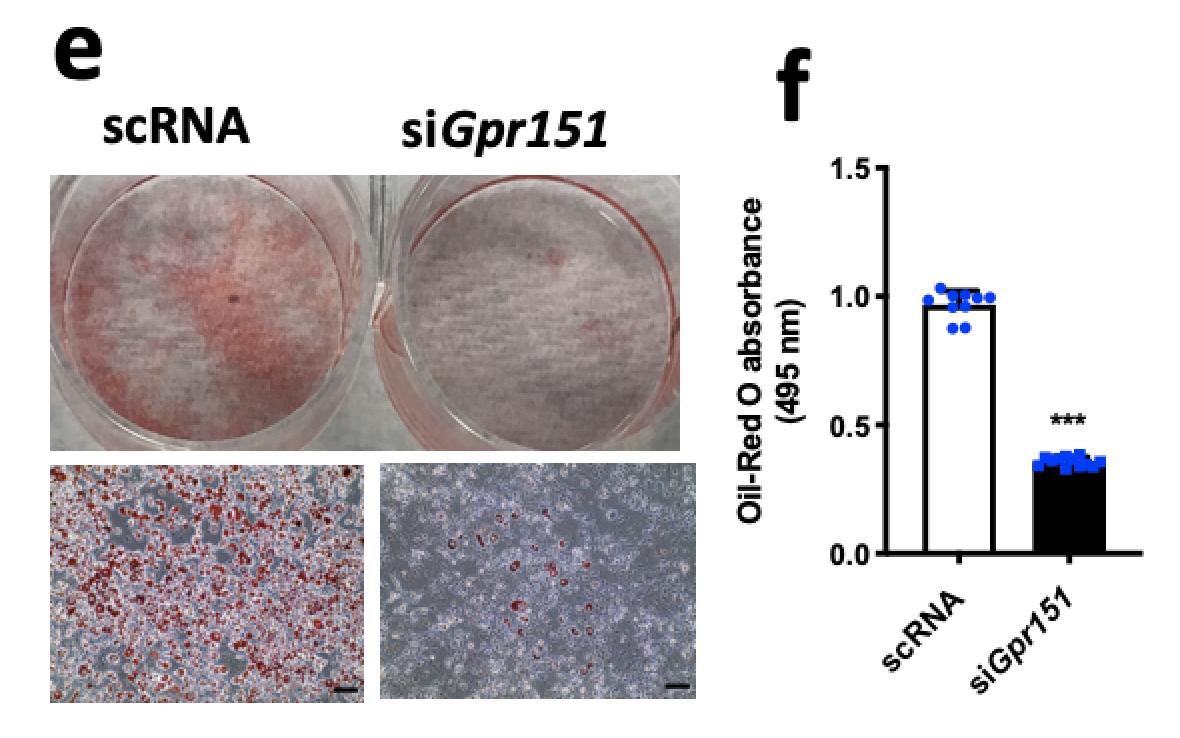
In the Rivas lab, we have several projects that extend the work presented in DeGAs.
DeGAs-PRS (dPRS): We propose dPRS, a method to enhance the interpretability of polygenic risk score (PRS) using DeGAs latent components.
Sparse reduced-rank regression (SRRR): In DeGAs, we took the summary statistics from univariate association scan across genetic variants and phenotypes. We propose a method to directly fit multi-response sparse regression models.
We provide a resource for the research community. We developed interactive DeGAs web application as a part of Global Biobank Engine, whose video tutorial is shown above.
The datasets used in the study are available from figshare.
Y. Tanigawa, and M. A. Rivas, Decomposed matrices used for the analysis described in ‘Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology’. https://doi.org/10.35092/yhjc.9202247.v1 (2019).