Medical relevance of protein-truncating variants across 337,205 individuals in the UK Biobank study
Apr 24, 2018
·
1 min read
Abstract
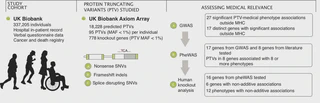
Protein-truncating variants can have profound effects on gene function and are critical for clinical genome interpretation and generating therapeutic hypotheses, but their relevance to medical phenotypes has not been systematically assessed. Here, we characterize the effect of 18,228 protein-truncating variants across 135 phenotypes from the UK Biobank and find 27 associations between medical phenotypes and protein-truncating variants in genes outside the major histocompatibility complex.
Protein-truncating variants can have profound effects on gene function and are critical
for clinical genome interpretation and generating therapeutic hypotheses, but their
relevance to medical phenotypes has not been systematically assessed. Here, we
characterize the effect of 18,228 protein-truncating variants across 135 phenotypes from
the UK Biobank and find 27 associations between medical phenotypes and
protein-truncating variants in genes outside the major histocompatibility complex. We
perform phenome-wide analyses and directly measure the effect in homozygous carriers,
commonly referred to as “human knockouts,” across medical phenotypes for genes
implicated as being protective against disease or associated with at least one phenotype
in our study. We find several genes with strong pleiotropic or non-additive effects. Our
results illustrate the importance of protein-truncating variants in a variety of
diseases.
Type
Publication
Published in Nature Communications, 2018
Using the UK Biobank population cohort, we investigated the genetic effects of Protein-truncating variants (PTVs) and the clinical impacts.